CardiacMotionFlow
CardiacMotionFlow is a Python-based tool for 2D cardiac MRI registration (or apparent flow generation), cardiac motion extraction, and cardiac pathology classification. Using deep learning, the method is both fast and lightweight. A package containing the code of the method, the pre-trained weights of the model, and the instructions for use can be downloaded here. The code of the method and the instructions for use are also available on this GitHub page. For more details about the method, the model, and the performance, please refer to the paper Explainable Cardiac Pathology Classification on Cine MRI with Motion Characterization by Semi-Supervised Learning of Apparent Flow.
CardiacSegmentationPropagation
CardiacSegmentationPropagation is a Python-based tool for cardiac image segmentation. Using deep learning with the spatial propagation of the segmentation, the method is both consistent and robust. A package containing the code of the method, the pre-trained weights of the model, and the instructions for use can be downloaded here. The code of the method and the instructions for use are also available on this GitHub page. For more details about the method, the model, and the performance, please refer to the paper 3D Consistent & Robust Segmentation of Cardiac Images by Deep Learning with Spatial Propagation.
Simul@trophy
Simul@trophy (available here) is a C++ based simulator tool that can simulate specified volume changes in different parts of the image. The tool was developed to generate synthetic time-series structural MRIs with specified ground truth atrophy. At its core, it implements a biophysical model of brain deformation with atrophy in Alzheimer’s disease. It can also simulate realistic variation of intensity.
medInria
medInria is a free medical imaging software developed at Inria. medInria aims at providing to clinicians state-of-the-art algorithms dedicated to medical image processing and visualization. Efforts have been made to simplify the user interface, while keeping high-level algorithms. medInria is available for Microsoft Windows, Linux Fedora / Ubuntu, Mac OS X, and is fully multi-threaded.
MUSIC
MUSIC: Multi-modality Platform for Specific Imaging in Cardiology
MUSIC is a software developed in collaboration with the IHU LIRYC in order to propose functionalities dedicated to cardiac interventional planning and guidance, based on the medInria software platform.
CardioViz3D
CardioViz3D targets research in cardiac imaging. It was funded by CardioSense3D project and aims at providing the researcher with a set of tools for pre-processing data and to visualize results of cardiac simulations. It has a robust visualization mechanism of any type of data, as well as a complete handling of time sequences of data.
SOFA
SOFA is an Open Source framework primarily targeted at real-time simulation, with an emphasis on medical simulation. It is mostly intended for the research community to help develop newer algorithms, but can also be used as an efficient prototyping tool.
LCClogDemons & SVF tools
LCClogDemons is an accurate and robust diffeomorphic registration framework based on the log-Demons.
It implements the symmetric Local Correlation Coefficient (LCC) as a similarity measure, and thus it is unbiased with respect to local linear intensity bias of the images.
The page also provide additional an ensemble of useful instruments, such a parallel transport with pole ladder, for working with stationary velocity fields.
ExoShape
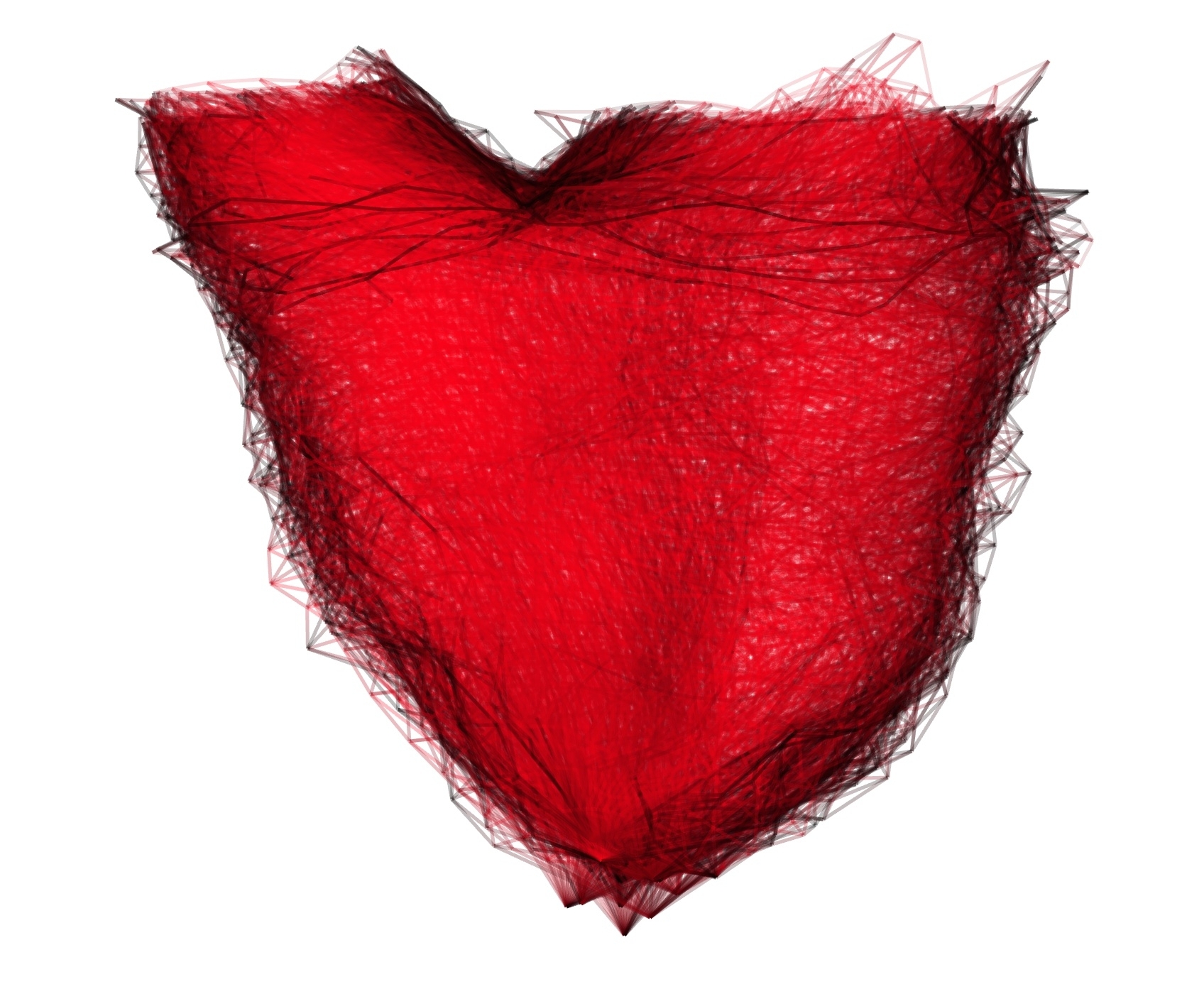

ExoShape is a toolbox to compute shape statistics
Registration Programming Interface (RPI)
A common Registration Programming Interface for image registration, in order to easily integrate different algorithms in a single software. Documentation is available.
Diffeomorphic Demons
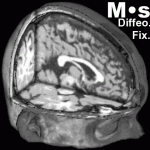
Diffeomorphic Demons is an efficient algorithm for the diffeomorphic registration of N dimensional images. It is based on Thirion’s demons algorithm but works on a Lie group structure on diffeomorphic transformations. Typical 3D medical images can be registered in less than three minutes on a 2 x 2.8 GHz quad-core Intel Xeon Apple Mac pro computer. Diffeomorphic demons is now included in MedINRIA‘s image fusion module. The source code has been integrated into ITK since version 3.8. A command-line software can be found on the Insight Journal. Additionally, some standalone binaries and tutorials may be found on the Stark Lab website.





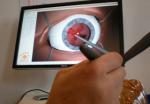
 Non invasive cardiac personalisation
Non invasive cardiac personalisation
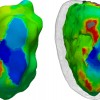 Simulation of ventricular tachycardia re-entry circuit
Simulation of ventricular tachycardia re-entry circuit
 heartMeshFine
heartMeshFine
 Electrophysiology
Electrophysiology
 Cardiac Fibres from in vivo Diffusion Tensor Imaging
Cardiac Fibres from in vivo Diffusion Tensor Imaging
