Personalization of Cardiac Models using Experimental Data and Machine Learning
Inria
- Research center(s) : CRI Sophia Antipolis – Méditerranée
- Inria Research team : ASCLEPIOS
- Principal investigator : Maxime Sermesant
- Field : Digital Health, Biology and Earth
- Theme : Computational Neuroscience and Medicine
International partner
- Area : North America
- Country : Canada
- Institution : University of Toronto
- Laboratory : Sunnybrook Research Institute
- Principal investigator : Mihaela Pop
Presentation of the associate team
Multi-scale computer modelling is a powerful tool that could be used to simulate in silico cardiac electrical activity and biomechanical function of individual heart. Imaging and 3D heart models built from images can help us understand the basis of structurally-diseased hearts at organ level and to predict in silico the changes in electro-mechanical function as a consequence of muscle remodelling in pathologic state (e.g. chronic infarction, a major cause of death). We hypothesize that MRI-based predictive models can help us identify new opportunities to intervene or to predict the outcome of ablation therapy, which currently has low clinical success. However, these predictive models need to be validated and thoroughly tested in preclinical experiments prior to their integration into the clinical stage. Hence, the next logical step for our joint Inria-SB efforts is to expand our experimental-theoretical framework and to personalize fast 3D heart models from in vivo MR-EP data. This translational step involves numerous challenging tasks from the modelling perspective since the in vivo imaging and physiological signals are rather noisy and obtained at a poor spatial resolution, potentially leading to erroneous customization of mathematical model parameters. However, this collaboration employs a rare combination of experiments and modelling specialists. Moreover, the originality of the proposed approach is to build upon machine-learning techniques rather than on data assimilation methods that are more explored in the literature but have inherent limitations (robustness to noise, local minima…).
Keywords: Associate teams CRI Sophia Antipolis – Méditerranée North America Canada
Abstract of the scientific program
Our aim is to develop high fidelity data-driven heart models using advanced cardiac MR imaging and computational methods. We propose an integrative 3-year collaborative program, by the end of which we hope to successfully validate these models and deliver personalized MRI-based electro-mechanical models for individual hearts for therapy planning and guidance.
Year 1: Develop anatomically and structurally accurate MR-based heart models. Several MR datasets have been acquired at SB in porcine hearts with chronic infarct. The main challenge here is to accurately integrate detailed structural information into the 3D meshes built from images and therefore to overcome the limitations of in vivo imaging.
Year 2: Optimization of inverse functions for the local personalization of various electrical and mechanical parameters using the 3D models built in O1. A particular attention will be given to the personalization of model parameters in the peri-infarct zone where arrhythmia foci resides.
Year 3: Assess the capacity of these models to predict the critical sites for ablation therapy and guide the intervention. We will validate the predictions with the actual measures of abnormal heart rhythms and the results from the performed ablation therapy.



 Non invasive cardiac personalisation
Non invasive cardiac personalisation
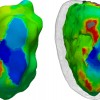 Simulation of ventricular tachycardia re-entry circuit
Simulation of ventricular tachycardia re-entry circuit
 heartMeshFine
heartMeshFine
 Electrophysiology
Electrophysiology
 Cardiac Fibres from in vivo Diffusion Tensor Imaging
Cardiac Fibres from in vivo Diffusion Tensor Imaging
