- Head of Computational Cardiology at Inria Epione, Université Côte d’Azur
- Head of Multimodal Data Science at IHU Liryc
- Chair of AI & Biophysics at 3IA Côte d’Azur
- Co-founder and scientific advisor of inHEART
- Co-founder of Therapixel
Research Projects
- Coordinator of the H2020 project SimCardioTest
- Coordinator of the EIT Health project inEurHeart
- Co-PI of the ERC Grant ECSTATIC
- PI for Inria in the European EraCoSysMed project PARIS
- Coordinator of The Digital Heart research program at UCA IDEX JEDI
- Scientific coordinator of the Software MUSIC at IHU Liryc
- Co-PI of the software platform medInria
Publications
- Inria website (with PDFs through HAL)
- Google Scholar
- Web of Science
- Microsoft Academic
- PubMed.
Short Bio
Maxime Sermesant is a permanent researcher at Inria, the French National Research Institute in Digital Sciences and Technology. His innovative research work combines biophysical and statistical modelling with clinical data. This is by nature a very collaborative and multi-disciplinary work at the intersection of academic, clinical and industrial environments. His research interests include biomedical image processing, organ modelling and machine learning. The integration of these areas open up possibilities in clinical data analysis for diagnosis, and in pathology simulation for therapy planning. He pioneered the development of patient-specific models of the heart. He received his Diploma in General Engineering from Ecole Centrale Paris, France in 1999, his MSc from Ecole Normale Superieure de Cachan, France in 1999, and his PhD in Control, Signal and Image Processing from the University of Nice – Sophia Antipolis, France in 2003. From June 2003 to December 2005, he was a Research Fellow with the Cardiac MR Research Group, Guy’s Hospital, King’s College London, UK and since 2005, he is a Research Scientist at Inria. He obtained a chair of AI & Biophysics at the 3IA Côte d’Azur AI Institute, and created the Multimodal Data Science team at IHU Liryc in 2019.
Previous Research Projects
- European EraCoSysMed project SysAFib: shape analysis for atrial fibrillation.
- MD Paedigree: Model-Driven Paediatric European Digital Repository.
- VP2HF: Virtual Physiological Human for Heart Failure.
- ERC MedYMA: Biophysical Modeling & Analysis of Dynamic Medical Images.
- Euheart: an integrated project on cardiac modeling and its application for improved diagnosis and therapy planning.
- Virtual Physiological Human: a network of excellence for collaborative investigation of the human body as a single complex system.
- Health e-Child: an integrated healthcare platform for European paediatrics.
Videos
CardioFunxion Winter School, Lyon, FR
VPH Summer School, Barcelona, ES
Tweets
Research Topics: Computer Models, Group-wise Statistics and Medical Image Processing
I first developed an electromechanical model of the heart which was usable as prior knowledge in image processing tasks. It was a generic model which could provide some physiological constraints but its parameters were not adjusted to the patient. Since then, my research focus followed two main axes. First I developed methods to personalise such an electromechanical model of the heart to the clinical data of a patient, in order to help diagnosis and therapy planning. Second, I integrated biophysical and statistical methods in order to be able to model the cardiac function at a group level.
All along these years, my research has gone back and forth between modelling and imaging, and the most recent work was in combining computational physiology and computational anatomy. This is a very exciting area that integrates two different ways of targeting patient-specific medicine. On one hand, computational physiology tries to build a computer model of the patient based on biophysical models of the human body. On the other hand, computational anatomy aims at statistical learning from healthy and pathological groupwise data, in order to evaluate a specific patient against the model.
These two research areas have undergone a tremendous progress over the last decade, which can be clearly seen by the number of related publications and conferences. Milestones were achieved in developing new methods for adjusting generic models to patient data and for computing statistics on complex objects like shapes and deformations. The parallel development of computational power and numerical strategies now opens new avenues for the application and integration of these results.
I really see the biophysical and statistical approaches as complementary, because independently they lack important features. For the biophysical approach, without a statistical analysis of the studied population it is often impossible to know which are the important phenomena to model, among the multi-scale multi-physics features of the cardiac function. On the other hand, once computed, a statistical model is very hard to interpret without some mechanistic insights on the phenomena observed, and biophysical models are a great tool to explore such mechanisms.




 Non invasive cardiac personalisation
Non invasive cardiac personalisation
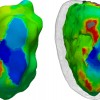 Simulation of ventricular tachycardia re-entry circuit
Simulation of ventricular tachycardia re-entry circuit
 heartMeshFine
heartMeshFine
 Electrophysiology
Electrophysiology
 Cardiac Fibres from in vivo Diffusion Tensor Imaging
Cardiac Fibres from in vivo Diffusion Tensor Imaging
