3D Anatomy Editor
Master Internship, Imagine team
Goal: To graphically edit anatomical models to match constraints
Contact: François Faure
Motivation
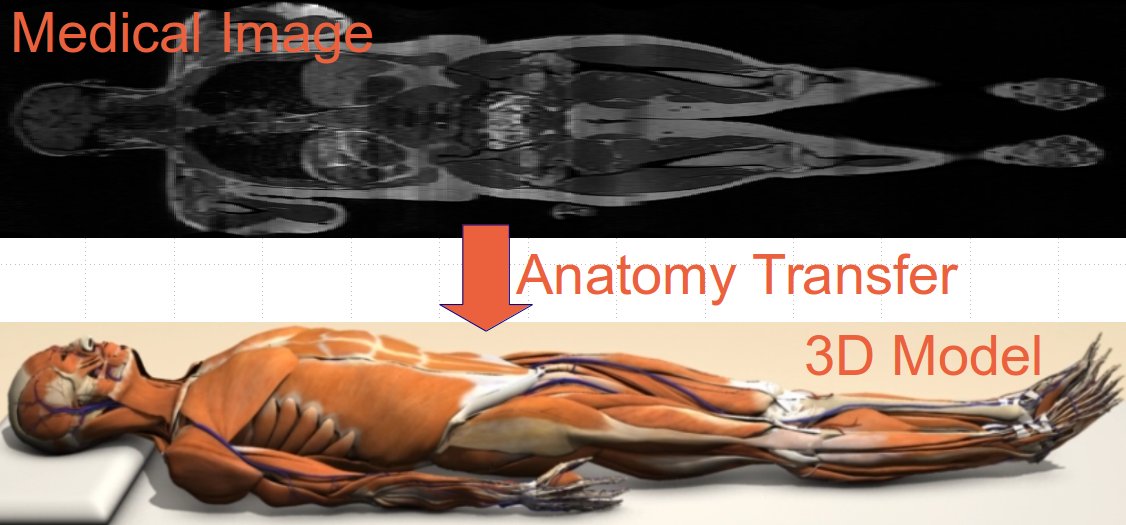
Using our recent work on Anatomy Transfer it is possible to automatically generate personalized 3D anatomical models based on medical images (MRI or CT) as illustrated below, by registrating a generic model to a personalized morphology.
However, while precision is surprisingly high when enough information is extracted from the image, the reconstructed surfaces do not perfectly match the image when few constraints are available, such as in the example below.
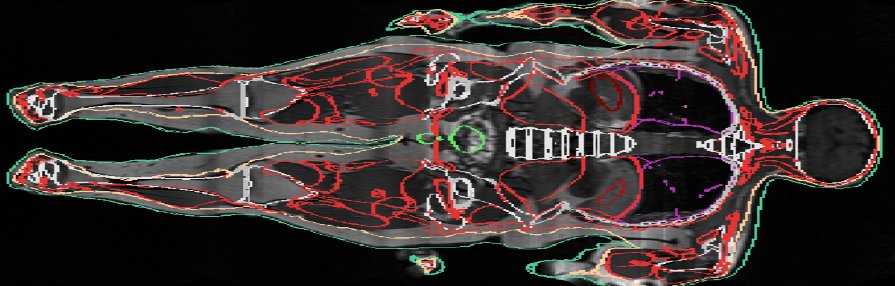
Model mismatch. In this example, only the skin was used to constrain the registration, and one can see in this slice that the transferred organs (in red and green) do not perfectly match the MRI image everywhere.
Proposed work
The objective of this project is to propose a novel graphical editor to quickly correct the mismatches.
In the first part, classical segmentation tools will be applied to locally correct the surfaces. Once the capabilities and limitations of this approach are clear, we will investigate higher level methods.
In the second part of this work, we will provide the user with novel tools to apply constraints in the registration process, by setting up registration constraints in the 2D slices or directly in 3D. These constraints will be converted to forces applied in the mechanical relaxation of a physical model.
Follow-ups
The results may be used to improve the technology of the Anatoscope startup company. This work may be continued as a PhD thesis, or engineering work in the company.
Profile
- Master student in Computer Science or Applied Mathematics.
- Programming skills in C++ or python or java are mandatory.
- Background in mathematics (especially linear algebra, geometry, and statistics).
- Prior knowledge in the areas of computer vision, computer graphics and embedded programming are relevant.


1 ping
… [Trackback]
[…] Read More Infos here: team.inria.fr/imagine/2014/10/04/internship-anatomy-edition/ […]