Participants:
Annabelle Collin, Clair Poignard, Olivier Saut
Clinicians involved:
J. Palussière, F. Cornelis, G. Kantor, H. Loiseau and H. Fathallah-Shaykh.
We have tested this strategy on metastasis to the lung from distant tumors (kidney, bladder, thyroid). We dispose of time series of CT-scans of patients that are only under monitoring (i.e. without treatment). We use only two CT-scans to parametrize our problem and try to recover the following ones.
Lung metastasis
The test case presented below concerns a metastasis to the lung of a bladder tumor. On the left are the 3 CT-scans that constitute our data set. No treatment was given to the patient during this period. We used only the image of June and September to perform the simulation.
In the middle plot, measured volumes of the metastasis (based on the scan) in time are the circle points while the continuous line is the volume predicted by the simulation. On the right images, the tumor given by the simulation for September and December is in red (top row). Other examples for metastasis to the lung are given in the publications.
- T. Colin, A. Iollo, D. Lombardi, O. Saut System Identification in Tumor Growth Modeling Using Semi-empirical Eigenfunctions. Math. Models Methods Appl. Sci. 22, 1250003 (2012).
- T. Colin, A. Iollo, D. Lombardi and O. Saut, Prediction of the Evolution of Thyroidal Lung Nodules Using a Mathematical Model, ERCIM News, No 82, pp. 37-38, July 2010.

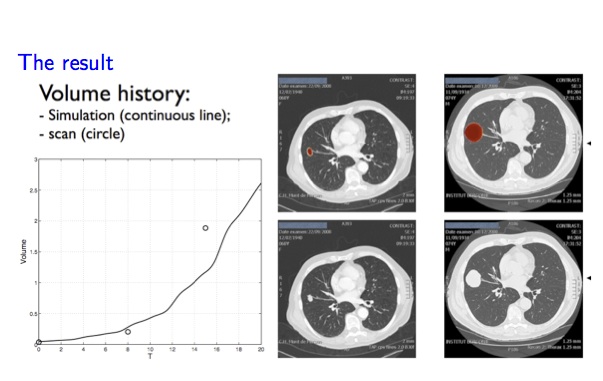
Lung metastasis growth prediction. Left: Data. Right: Model simulations and predictions.
Liver metastasis
We have also some preliminary results concerning metastasis to the lung of a GIST (Gastro-Intestinal Stromal Tumor). When the metastasis is discovered, the patient receives a targeted therapy called Imatinib until he escapes the treatment. He then receives a second therapy with Sunitinib until the next escape time. In the example below we show on the left the clinical data: time curve of the volume of the metastasis measured on the successive CT-scans and the CT-scans corresponding (from left to right and from top to bottom) to the control by the first treatment, then the escape and then the control but the second treatment. Right images is a fit of the model to the volume data.
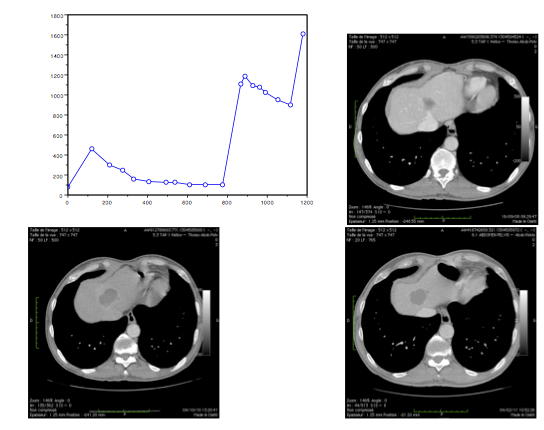

Liver metastasis. Left: Data provided by the Institut Bergonié. Right: Model fit.
Meningioma
Glioblastoma (GBM)
With H. Fathallah (University of Alabama at Birmingham), we have developed a 3D model of glioblastoma expansion that shows the three layer structure of GBM: a necrotic core, a proliferative rim surrounded by a cloud of invasive cells. These elements can be seen on different sequences of MRI (T1, T1 gado and Flair). It is still an open problem to characterize precisely these elements on these images. Below is shown a simulation of GBM expansion together with an MRI and a biopsy of a glioblastoma.
- Th. Colin, A. Iollo, J.-B. Lagaert and O. Saut, An inverse problem for the the recovery of the vascularization of a tumor.
- Th. Colin, H. Fathallah, J.-B. Lagaert, O. Saut, A Multilayer Model for GBM: Effects of Invasive Cells and Anti-Angiogenesis on Growth. Submitted.
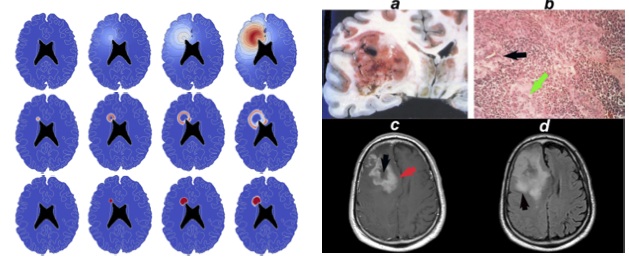
Glioblastoma. Left: Model simulations. Right: Data provided by the University of Alabama.

