WellReader is a MATLAB application developed in the Ibis group. The software implements computational methods developed for the analysis of the dynamics of gene expression in microorganisms, measured by means of fluorescent and luminescent reporter genes.
As input, WellReader reads the primary data file produced by a 96-well microplate reader, containing the time-series measurements of the absorbance (optical density) levels as well as the fluorescence and luminescence levels in each well (if available).
The main view mimics the microplate, and various modules exist to analyze the data, in particular to detect outliers, correct data by background substraction, smooth the data, compute promoter activities and concentrations, compare expression profiles, etc.
At each step of the analysis, the results of the actions on the data are stored and can be saved and reloaded for further analysis.
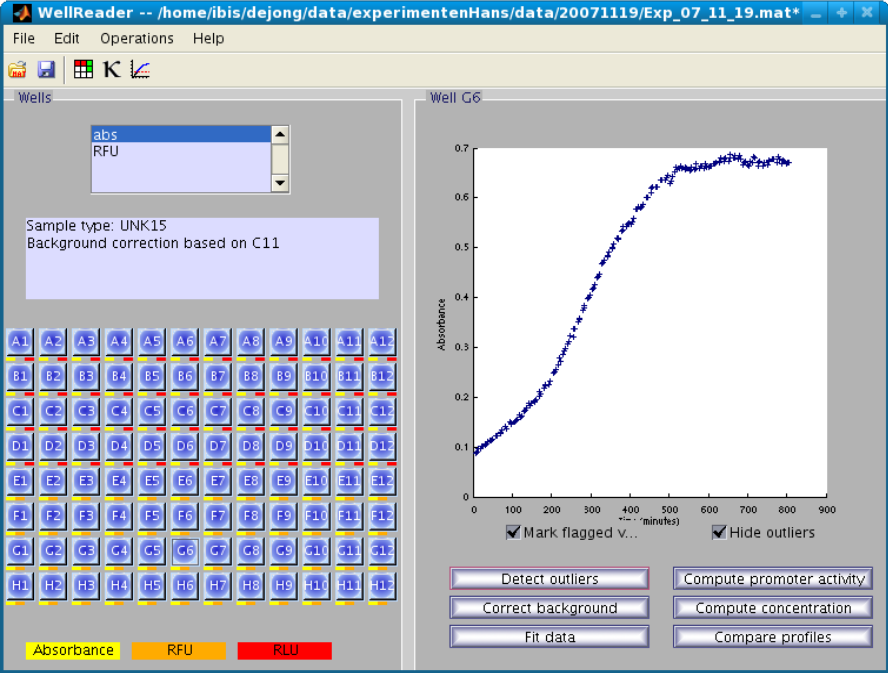
Screenshot of WellReader
Obtaining and using WellReader
WellReader is freely available under an LGPL licence. The program is written in MATLAB and is available both as source code (M files) and (on request) as compiled code (platform-specific binary files).
More information on how to use WellReader can be found in the tutorial and the user guide on the WellReader web site.
Developers of WellReader
The main developers of WellReader are Guillaume Baptist, Bruno Besson, Frédéric Boyer, Hidde de Jong, Hans Geiselmann, Jérôme Izard, and Delphine Ropers,.
References
- F. Boyer, B. Besson, G. Baptist, J. Izard, C. Pinel, D. Ropers, J. Geiselmann, H. de Jong. WellReader: A MATLAB program for the analysis of fluorescence and luminescence reporter gene data. Bioinformatics, 26(9):1262-1263, 2010
- H. de Jong, C. Ranquet, D. Ropers, C. Pinel, J. Geiselmann. Experimental and computational validation of models of fluorescent and luminescent reporter genes in bacteria. BMC Systems Biology, 4:55, 2010


