WellInverter is a web application implementing linear inversion methods for the reconstruction of gene expression profiles from fluorescent or luminescent reporter gene data.
As input, WellInverter reads the primary data file produced by a multi-well microplate reader, containing time-series measurements of the absorbance (optical density) as well as the fluorescence and luminescence levels in each well (if available). Various modules exist to analyze the data, in particular for detecting outliers, subtracting background, estimating growth rates, promoter activities and protein concentrations, visualizing expression profiles, computing statistics on replicate profiles, etc.
More information on how to use WellInverter can be found in the tutorial.
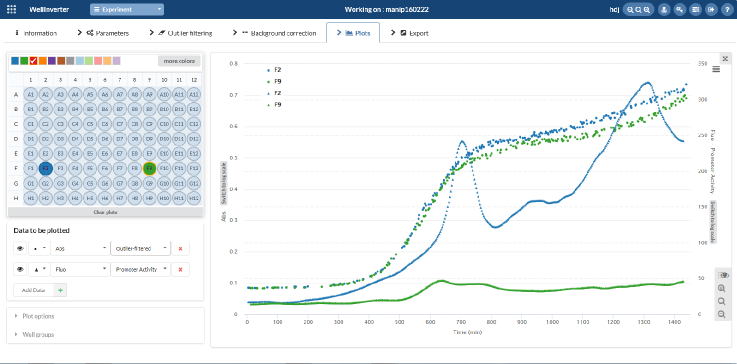
Screenshot of WellInverter
WellInverter architecture
The server part of WellInverter is based on the Python library WellFARE, the computational core of the application. It also provides methods for managing experimental and user data as well as storing analysis results in JavaScript Object Notation (JSON) format.
The client part of WellInverter is the graphical user interface of the application, working in a web browser. It allows the user to upload, analyze, and visualize the results of a reporter gene experiment as well as downloading the results for further treatment. The client part is written in Javascript, and communicates with the server using JSON-encoded data and Ajax (Asynchronous JavaScript) calls.
WellInverter uses a plug-in system for defining high-level routines for parsing data files produced by microplate readers from different manufacturers. Parsers for Tecan and Perkin-Elmer microplate readers are currently available.
Access to WellInverter
WellInverter demo. A demo version of WellInverter can be accessed at the following address:
https://wellinverter.inrialpes.fr
The demo version does not allow the user to upload his/her own data sets. However, sample data is supplied: two primary data sets of E. coli fluorescent reporter gene experiments, as well as a completely treated version of one of these data sets.
WellInverter Inria. WellInverter is deployed on an Inria server:
http://ibis-public.inrialpes.fr:8000/
No user registration is needed, but the absence of a user account does not allow treated data files to be stored on the server and the user needs to download and locally save the analysis results before the end of the session.
WellInverter stand-alone. Users wishing to locally install WellInverter on their computer can download the stand-alone version from the following address:
https://gitlab.inria.fr/WellInverter/WellInverter_Standalone_Windows
The installer is password-protected, please contact Hidde de Jong for obtaining the password.
The Python library WellFARE, implementing the linear inversion methods on which WellInverter is based, is separately available under an LGPL license at the following address :
https://github.com/ibis-inria/wellfare
Developers of WellInverter
WellInverter has been developed by Yannick Martin, Michel Page, Valentin Zulkower, and Hidde de Jong, with help from Johannes Geiselmann and Delphine Ropers. The development of the application has been supported by the French Institute for Bioinformatics (IFB), in particular Christophe Blanchet.
References
- V. Zulkower, M. Page, D. Ropers, J. Geiselmann, H. de Jong. Robust reconstruction of gene expression profiles from reporter gene data using linear inversion. Bioinformatics, 31(12):i71-i79, 2015
- Y. Martin, M. Page, C. Blanchet, H. de Jong. WellInverter: A web application for the analysis of fluorescent reporter gene data. BMC Bioinformatics, 20(1):309, 2019


